Data Exchange Formats and Metadata Standards for Research Data Management and Sharing
FISH-Omics Format for Chromatin Tracing (FOF-CT)
A key output of the (4D Nucleome) project is the open release of datasets that measure the spatial arrangement of DNA, RNA, and proteins within the human cell nucleus, thereby uncovering the functional dynamics of the genome in three dimensions and over time (referred to as 4D).
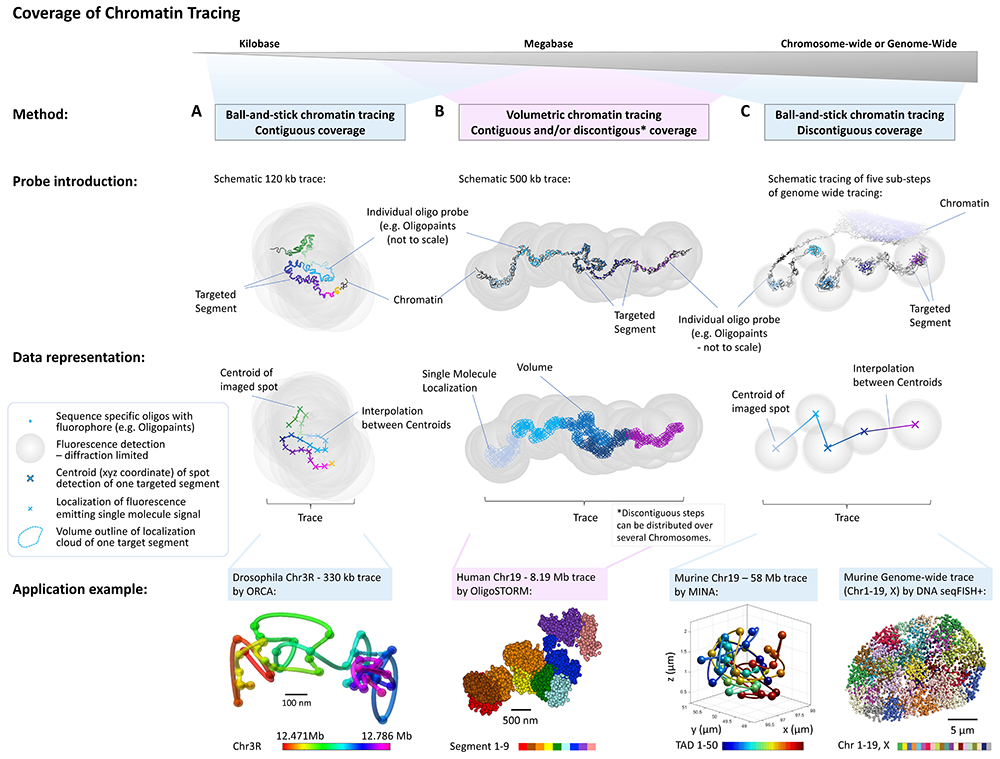
FISH omics methods can be utilized to map chromatin structures across multiple genomic scales (Figure credit: Sarah Aufmkolk; Dekker et al. 2023, Mol.Cell.)
FISH Omics (a.k.a. Multiplexed FISH) methods provide an expanded understanding of how higher-order chromosome structure relates to transcriptional activity and cell development.
Various protocols have been developed in the past few years, which can be divided into two main categories: ball-and-stick or volumetric Chromatin Tracing (for a review, see Dekker et al. 2023).
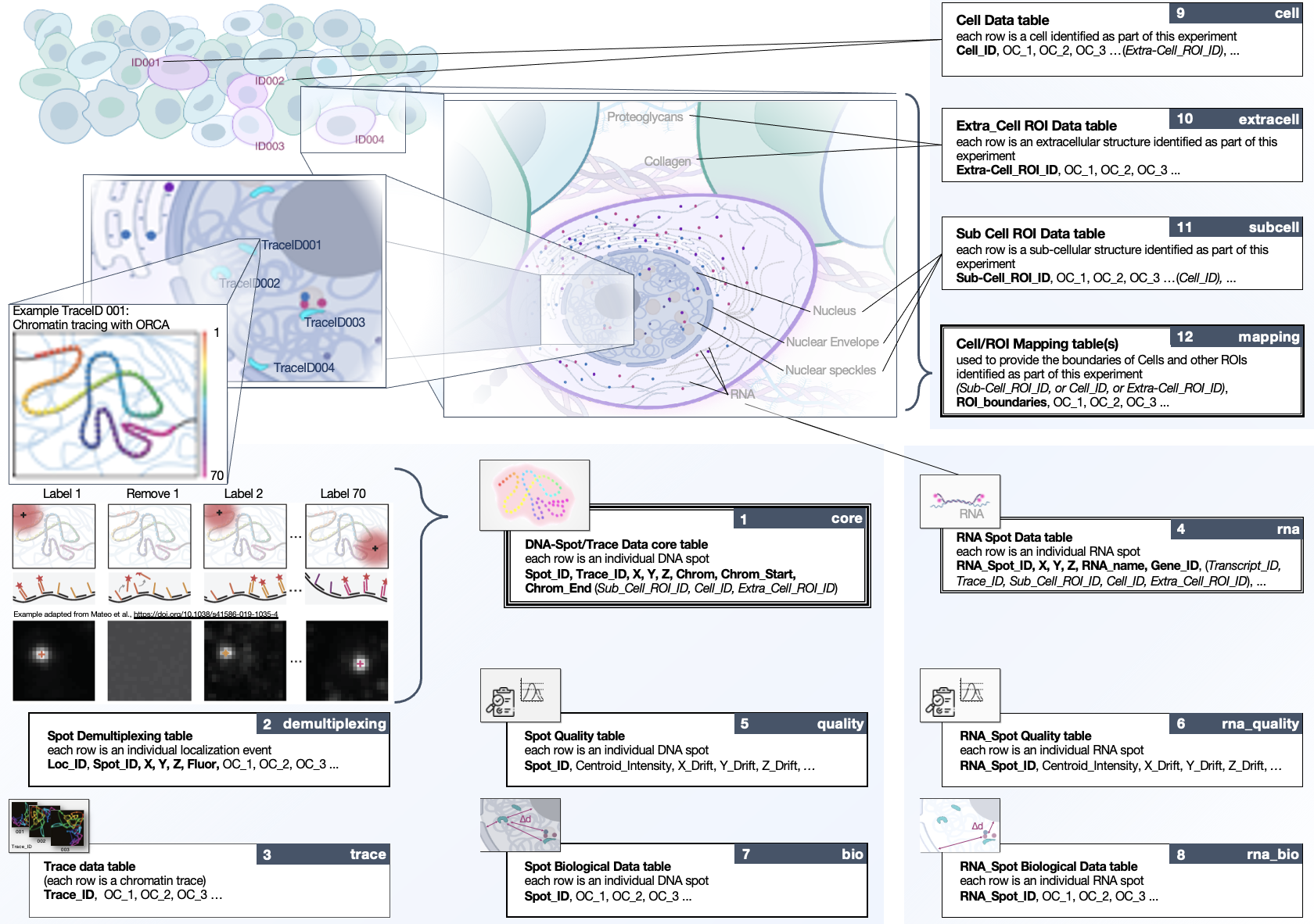
The 4DN FISH Omics Format - Chromatin Tracing (FOF-CT) is directly compatible with several ball-and-stick FISH-omics techniques including, but not limited to, Optical Reconstruction of Chromatin Architecture (ORCA), Multiplexed Imaging of Nucleome Architectures (MINA), DNA Multiplexed error-robust fluorescence in situ hybridization (DNA-MERFISH), and In-situ Genomic Sequencing (IGS).
In addition, the format is designed to be consistent with the ongoing development of extensions that will encompass volumetric chromatin tracing methods, such as OligoSTORM and OligoDNA-PAINT (Belivau et al. 2017, Bintu et al. 2018, Nir et al. 2018, Luppino et al. 2020).