Data Exchange Formats and Metadata Standards for Research Data Management and Sharing
NBO-Q Microscopy Metadata Specifications
Light microscopy is a powerful tool for quantitative image analysis in biomedical research. It is used to probe the real-time dynamics of sub-cellular structures. Community-agreed standards for image metadata are crucial to ensure the correct interpretation, reproducibility, and reuse of image data. For almost 20 years, the Open Microscopy Environment (OME) served this purpose for the community by developing and maintaining the OME Data Model, which is implemented in OME-TIFF, Bio-Formats, and OMERO.
Over that same period, imaging scientists developed new imaging modalities and found creative ways to use imaging in their research, which were unanticipated by the original model developers. Gaps between the model and image metadata necessitated revising and expanding the model to improve metadata capture practices and provide a flexible foundation for future model modification in response to further developments in imaging technology.
In 2021, the NIH-funded 4D Nucleome program (4DN), BioImaging North America (BINA), QUAREP-LiMi, and the Open Microscopy Environment (OME) released a revision of the Core OME data model called 4DN-BINA-OME-QUAREP (NBO-Q) Microscopy Metadata model (Hammer et al., 2021. Nature Methods).
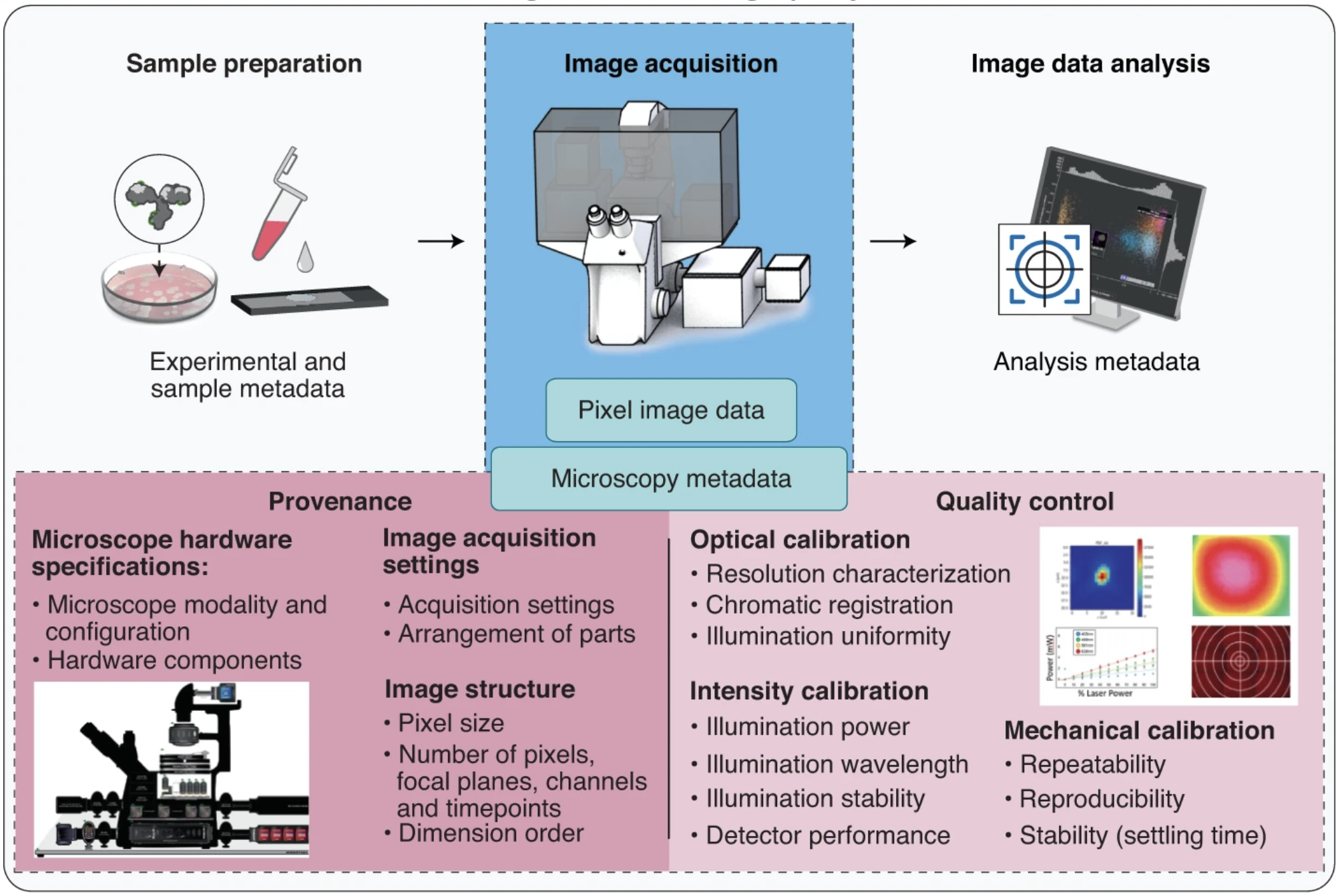
NBO-Q is a Tiered system of metadata specifications. It includes three extensions to capture instrument hardware specifications and image acquisition metadata from experiments performed on transmitted and wide-field fluorescence microscopes (Basic extension) and confocal and other advanced fluorescence imaging modalities (Advanced and Confocal extension). In addition, the Calibration and Performance extension provides a framework for managing microscope quality control metrics and processes.