Software tools for Research Data Management and Sharing
OMEGA: Open Microscopy Environment Integrated Analysis
Dissecting the interplay between HIV-1 and human cells during viral entry
After HIV-1 fuses with a target cell membrane, the virion core is delivered into the cytosol of the infected cell. A DNA copy (cDNA) of the HIV-1 RNA genome is produced by the HIV-1 reverse transcriptase (RT), and the cDNA is ligated to host cell chromosomal DNA by HIV-1 integrase.
Quantitative analysis of viral trajectories gives insight into viral-host interactions
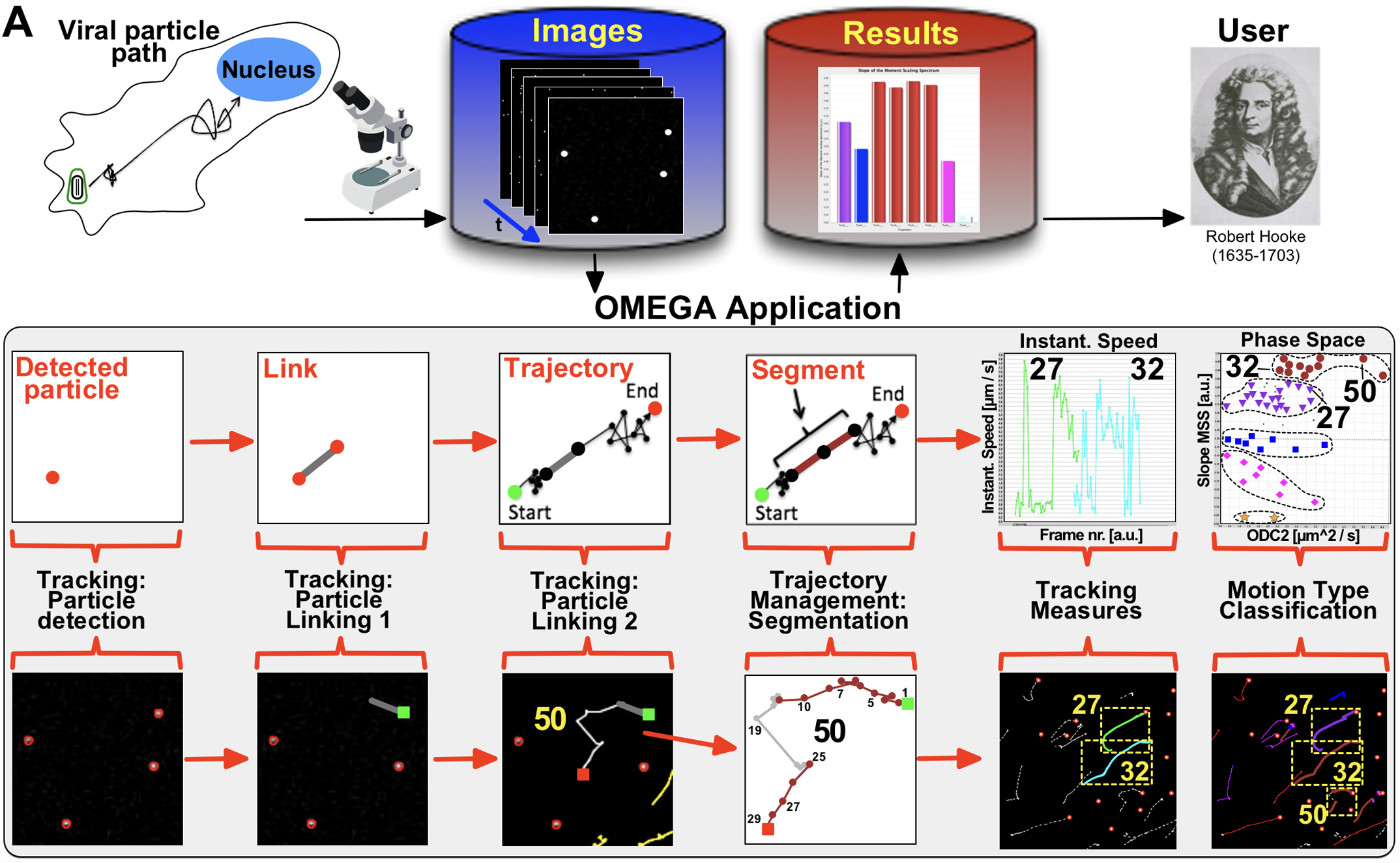
The complex interplay between HIV-1 and cellular components can be solved using dynamic imaging followed by rigorous computational analysis of viral particle trajectories. A major hurdle has been the difficulty of keeping track of multiple moving viral particles with sufficient temporal and spatial resolution while probing multiple experimental contexts.
A three-pronged multidisciplinary approach
To meet this challenge, the Strambio De Castillia group is engaged in a multidisciplinary, collaborative effort to develop integrated workflows that will provide the analysis, data management, and visualization capabilities required to drive a complete understanding of viral-host interactions in the context of living human tissues. The project articulates on three fronts:
- Real-time recording of HIV-1 viral core movement in infected human cells.
- Viral particle tracking and motion analysis.
- Building a novel bio-image informatics framework, called OMEGA, to support the comprehensive examination of particle movement across multiple experimental models and conditions.
OMEGA project output
- Rigano A, Galli V, Clark JM, Pereira LE, Grossi L, Luban J, et al. OMEGA: a software tool for the management, analysis, and dissemination of intracellular trafficking data that incorporates motion type classification and quality control. bioRxiv 2018. Available from: http://dx.doi.org/10.1101/251850.
- Rigano A, Galli V, Gonciarz K, Sbalzarini IF, Strambio-De-Castillia C. An algorithm-centric Monte Carlo method to empirically quantify motion-type estimation uncertainty in single-particle tracking. bioRxiv 2018. Available from: http://dx.doi.org/10.1101/379255.
- Rigano A, Strambio-De-Castillia C. Proposal for minimum information guidelines to report and reproduce results of particle tracking and motion analysis. bioRxiv 2017. Available from: http://dx.doi.org/10.1101/155036.
